 2017.ĭong W, Liu J, Yu J, Wang L, Zhou S. NOVOPlasty: de novo assembly of organelle genomes from whole genome data. A platform for efficient genotyping in Musa using microsatellite markers. 2003.Ĭhristelová P, Valárik M, Hřibová E, Channelière S, Roux N, Doležel J. NISC comparative sequencing program LAGAN and Multi-LAGAN: efficient tools for large-scale multiple alignment of genomic DN. 2014.īrudno M, Do CB, Cooper GM, Kim MF, Davydov E, Green ED, Sidow A, Batzoglou S. Trimmomatic: a flexible trimmer for Illumina sequence data. Circular chloroplast chromosomes: the grand illusion. MISA-web: a web server for microsatellite prediction. 
2010.īeier S, Thiel T, Münch T, Scholz U, Mascher M. 
FastQC: a quality control tool for high throughput sequence data. IRscope: an online program to visualize the junction sites of chloroplast genomes. Phylogenomic and comparative analyses of Coffeeae alliance (Rubiaceae): deep insights into phylogenetic relationships and plastome evolution. The plastome data generated in the study would provide the baseline data for future phylogenetic studies.Īmenu SG, Wei N, Wu L, Oyebanji O, Hu G, Zhou Y, Wang Q. indandamanensis in section Musa as a sister to M. The current phylogenomic study confirmed the placement of M. It also possesses 30, 9, 7, 9, 4, 2, and 7, mono, di, tri, tetra, penta, hexa, and compound SSRs, respectively. Corroborating the previous studies, some nucleotide hotspot regions ( atpH– atpI, ndhG– ndhI, and rpoB– petN and psbC– psaB) were also revealed during this comparative study. Moreover, it also depicted 21, 29, 11, and 3, forward, palindromic, reverse and complement repeats, respectively. indandamanensis is 170,207 bp long in size and depicts a typical quadripartite structure having 88,753pb long LSC, 10,820 bp long SSC, and two 35,317 bp long IRs. kattuvazhana and different taxa belonging to the section Musa. indandamanensis and compared it with closely allied species M. In the present study, we sequenced and characterized the complete plastome of M. One of the economically important species, Musa indandamanensis, also known as a sweet banana with orange pulp, is confined to the Andaman and Nicobar Islands (A&N) in India. Many species of this genus are narrowly endemic to India and are used as a food source or herbal medicines by the tribal communities. To export a folder from the Local, simply select the folder and in the Toolbar click on “Export”, “Export Folder…”, choose the file destination and name, and click “Save”.Musa is one of the most economically important genera distributed throughout the globe. Fill in as much information as possible for future reference.Ī very convenient tool is the ability to export and import folders of primers. Here, various metadata such as Gene, Organism, Direction etc. Highlight the primer in the Document Table, and go to the “Info” tab in the Document Viewer. Once a primer has been created, any associated information can be edited. Here, enter the primer sequence, name, and in the “Type” dropdown menu select “Primer”. To add a new primer, highlight the destination folder, then in the Menu Bar select “Sequence” followed by “New Sequence” from the dropdown menu. From the Toolbar, click “Add” then “New folder…” and enter the new folder name. 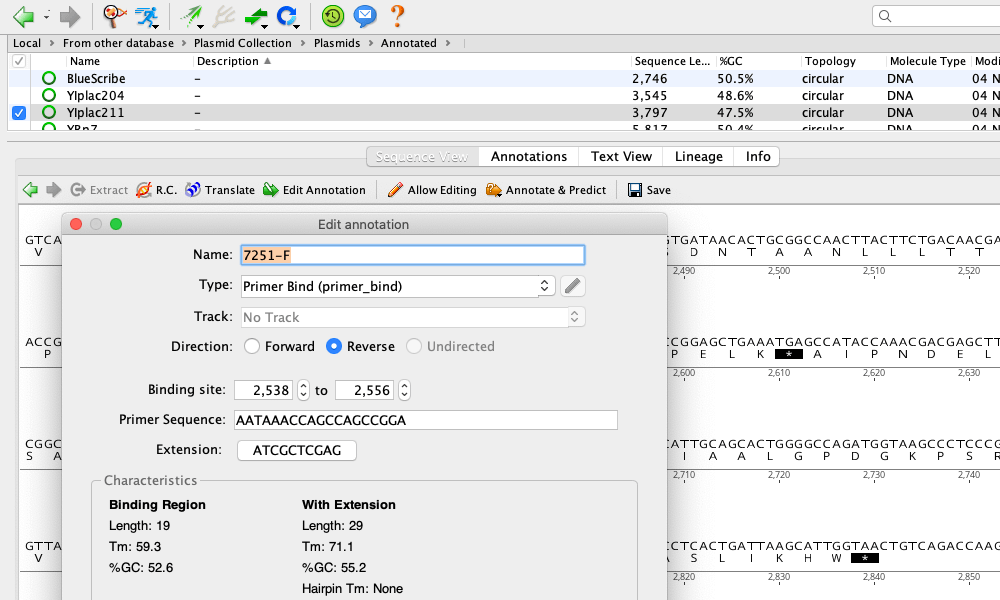
To create a new folder in the directory, highlight the local folder in the Sources Panel. If a primer folder does not exist in the Local Directory, one should be made. SI Barcode Network Informatics Documentation
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
